|
I am a Senior Scientist in Machine Learning in the Data AI and Genomics Group at Merck Research Laboratories. My research focuses on pretraining large foundation models, optimizing training through efficient kernels, and evaluating model performance. My broader research interests include foundation models, deep learning, machine learning systems, training optimization, and explainable AI. I received my Ph.D. in Computer Science from Rensselaer Polytechnic Institute (2025), where I was advised by Professor James A. Hendler and Professor Pingkun Yan. I also hold a M.S. in Computer Science from Rensselaer Polytechnic Institute, a M.S. in Biomedical Engineering from Johns Hopkins University, and a B.S. in Electrical Engineering from PES Institute of Technology. I have been advised by wonderful mentors: Professor Sridevi Sarma at JHU, Dr. Qiang Chen and Dr. Kristen Maynard at LIBD, Professor Carl Petersen at EPFL, and Professor Achuta Kadambi at MIT.Email / CV / Resume / Google Scholar / Twitter / LinkedIn / GitHub |
|
|
|
|
|
|
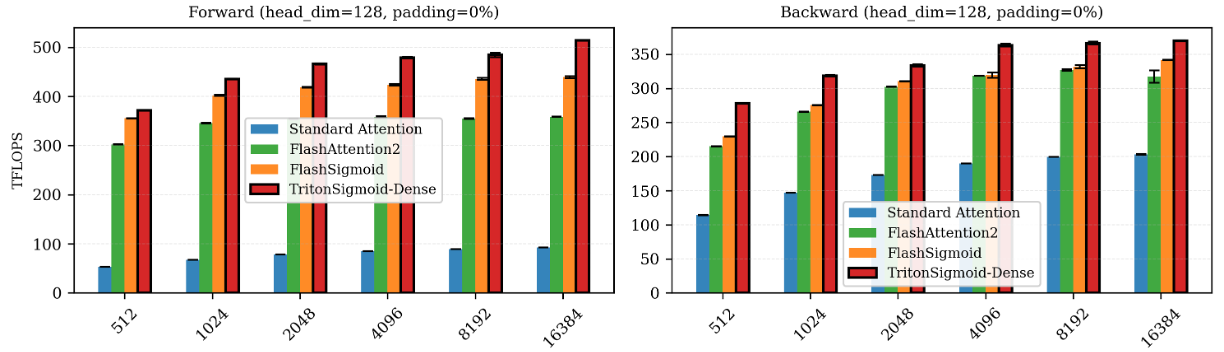
Vijay Sadashivaiah, Georgios Dasoulas, Judith Mueller, Soumya Ghosh arXiv preprint, 2026 paper / code We demonstrate that replacing softmax with sigmoid attention in biological foundation models yields 25% improved cell-type separation, up to 10% faster training, and improved stability. We develop TritonSigmoid, an efficient GPU kernel achieving 515 TFLOPS on H100s. |
|
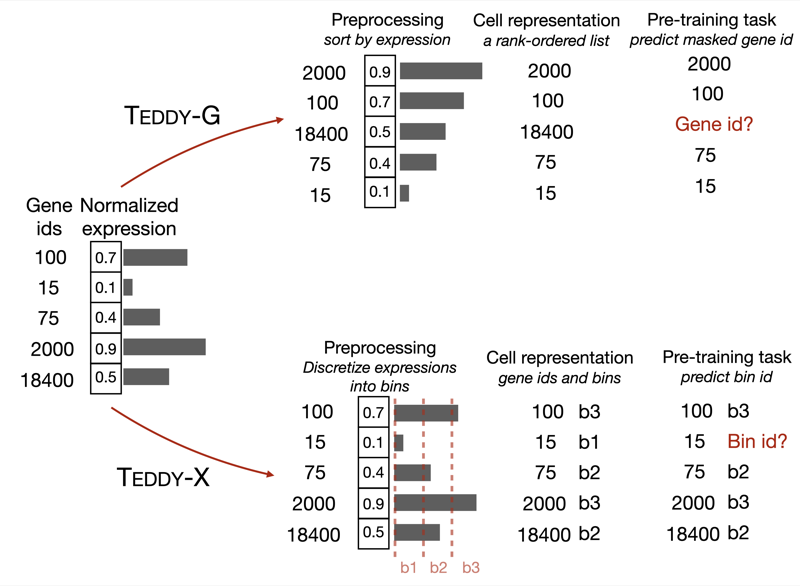
Alexis Chevalier, Soumya Ghosh, Urvi Awasthi, James Watkins, Julia Bieniewska, Nichita Mitrea, Olga Kotova, Kirill Shkura, Andrew Noble, Michael Steinbaugh, Vijay Sadashivaiah, George Dasoulas, Julien Delile, Christoph Meier, Leonid Zhukov, Iya Khalil, Srayanta Mukherjee, Judith Mueller GenBio Workshop at ICML, 2025 paper We present TEDDY, a family of transformer-based foundation models for single-cell biology, scaled to 116 million cells. The models (70M, 160M, and 400M parameters) can identify disease states and distinguish diseased from healthy cells, with performance improving predictably with data volume and parameter count. |
|
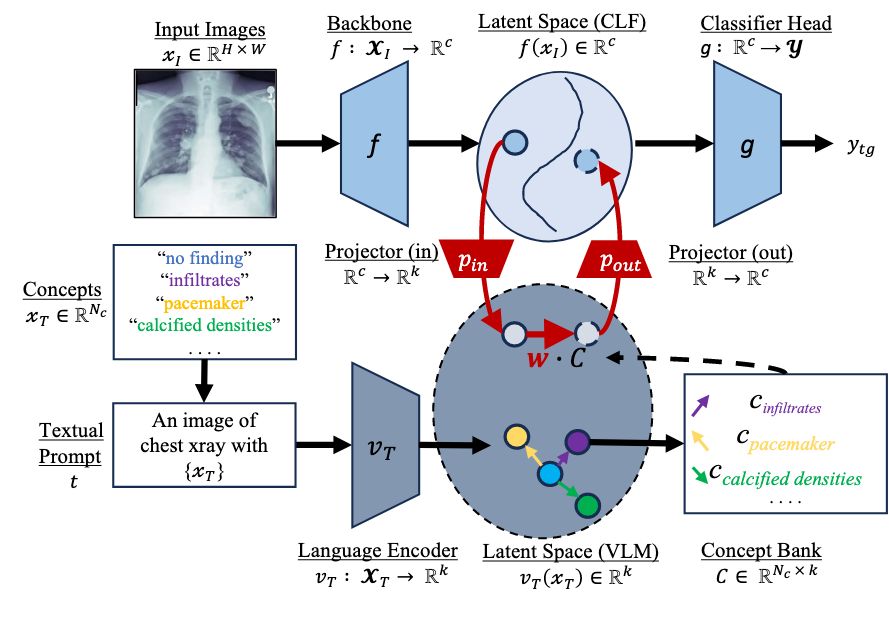
Vijay Sadashivaiah, Mannudeep K Kalra, Pingkun Yan, James A. Hendler AIM-FM at NeurIPS, 2024 paper We propose to explain chest X-ray pathology models using textual concepts. This is achieved by leveraging the joint latent space of image and text in vision-language models. |

|
Fnu Mohbat, Vijay Sadashivaiah, Keerthiram Murugesan, Amit Dhurandhar, Ronny Luss, Pin-Yu Chen TrustNLP at NAACL, 2024 paper We evaluated the influence of bias on multimodal text generation models. In particular, we studied the impact of visual augmentation using state-of-the-art diffusion models when generating text. |
|
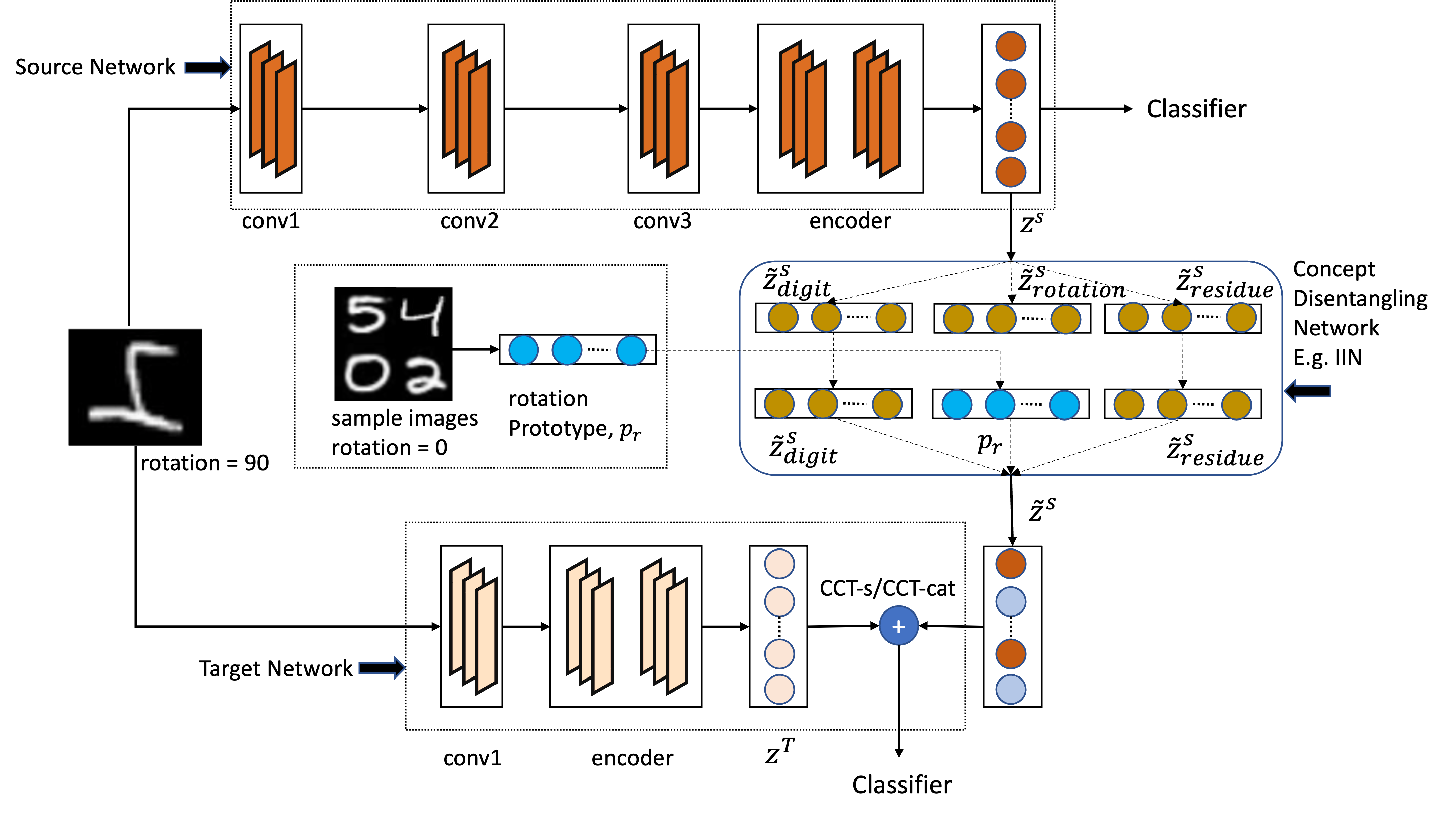
Vijay Sadashivaiah*, Keerthiram Murugesan*, Ronny Luss, Pin-Yu Chen, Christopher R. Sims, James A. Hendler, Amit Dhurandhar (* equal contribution) Transactions on Machine Learning Research (TMLR), 2024 paper We propose to suppress user-determined semantically meaningful concepts (viz. eyeglasses, smiling) from intermediate representations in computer vision tasks. |

|
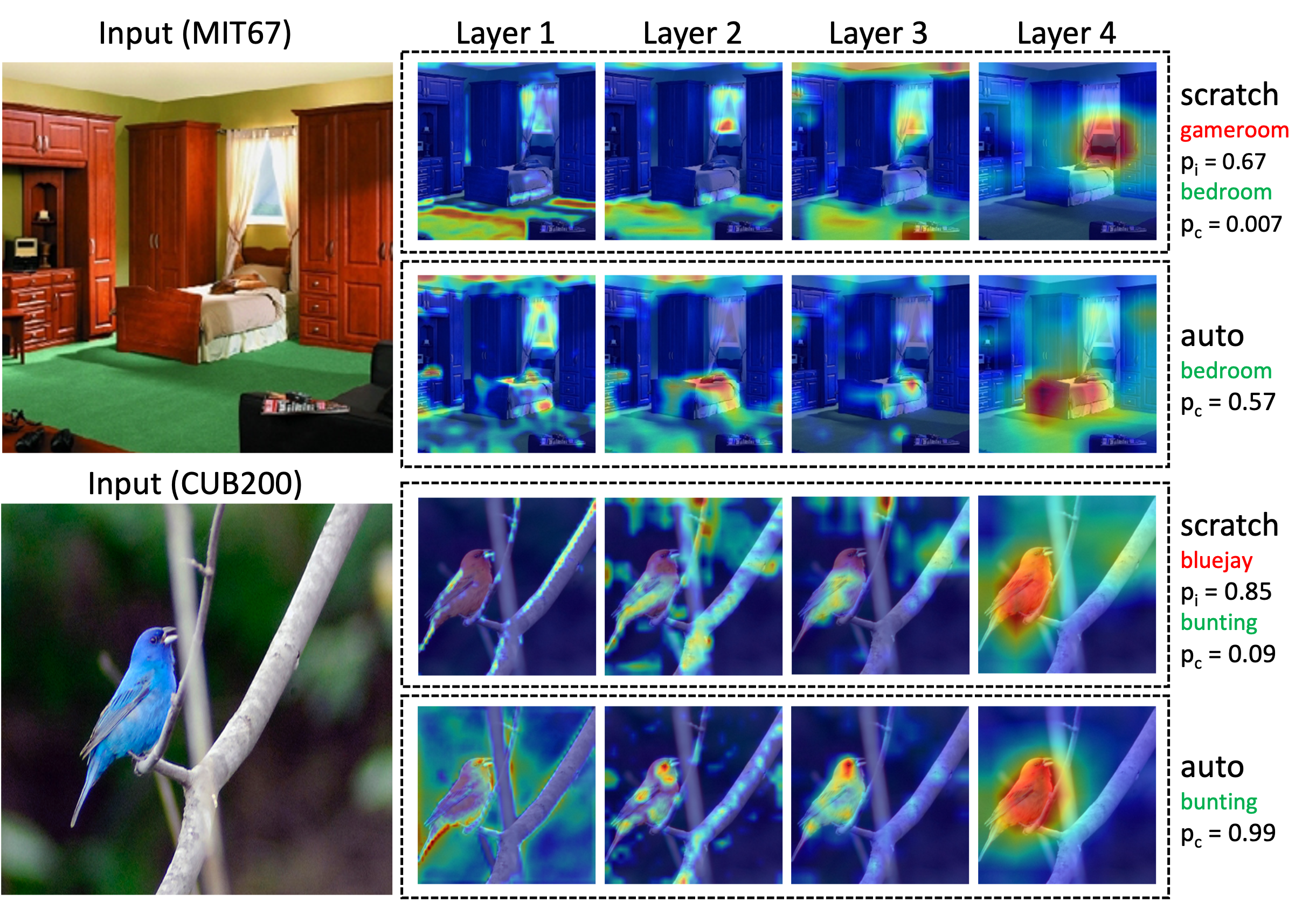
Keerthiram Murugesan*, Vijay Sadashivaiah*, Ronny Luss, Karthikeyan Shanmugam, Pin-Yu Chen, Amit Dhurandhar (* equal contribution) International Conference on Learning Representations (ICLR), 2022 paper / code We introduce multi-armed bandit based representation routing to improve transfer learning in computer vision tasks. |
|
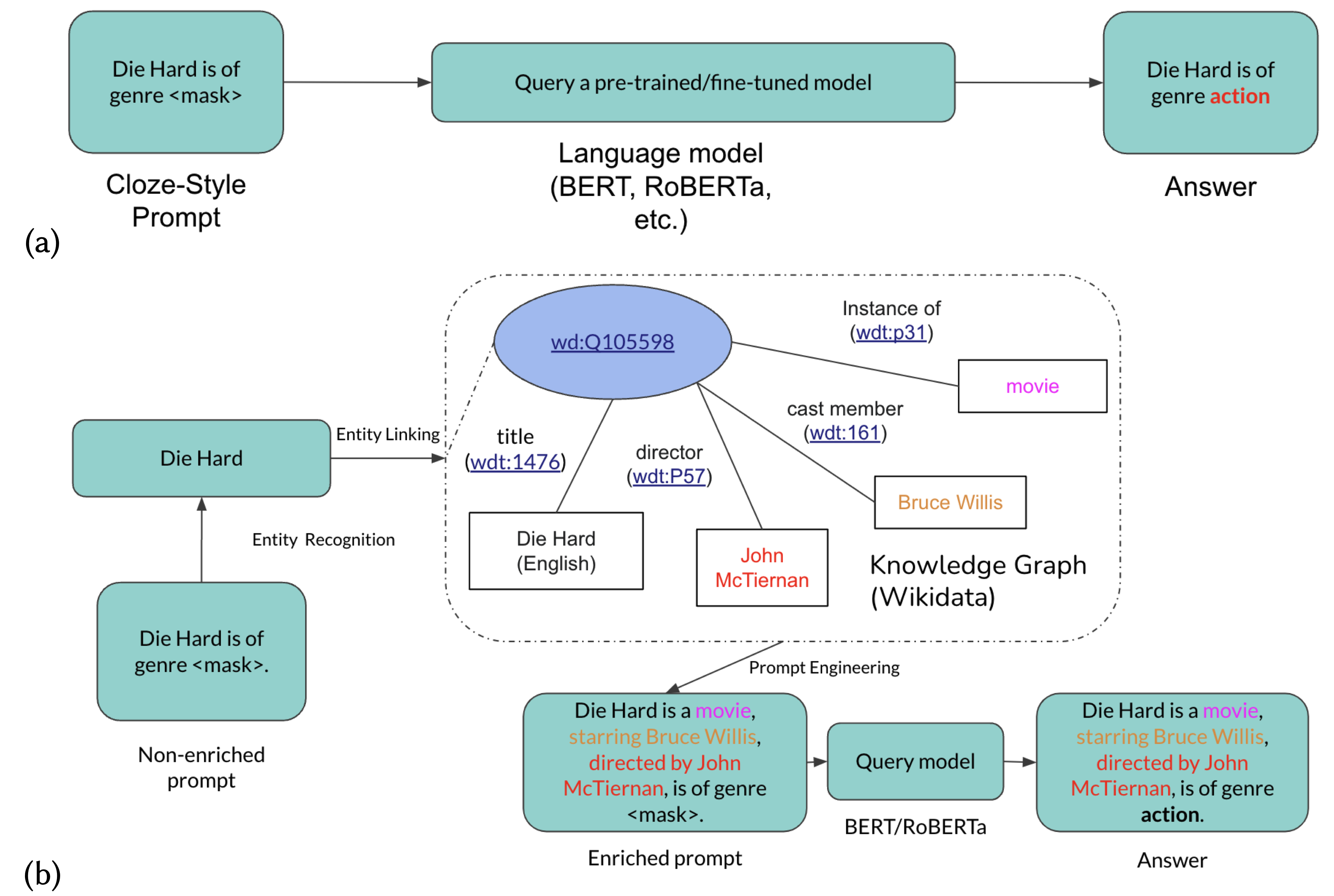
Ryan Brate, Minh-Hoang Dang, Fabian Hoppe, Yuan He, Albert Meroño-Peñuelar, Vijay Sadashivaiah Deep Learning for Knowledge Graphs Workshop at ISWC, 2022 paper We propose to imporve language model predictions by enriching the prompts from knowledge graphs. |
|
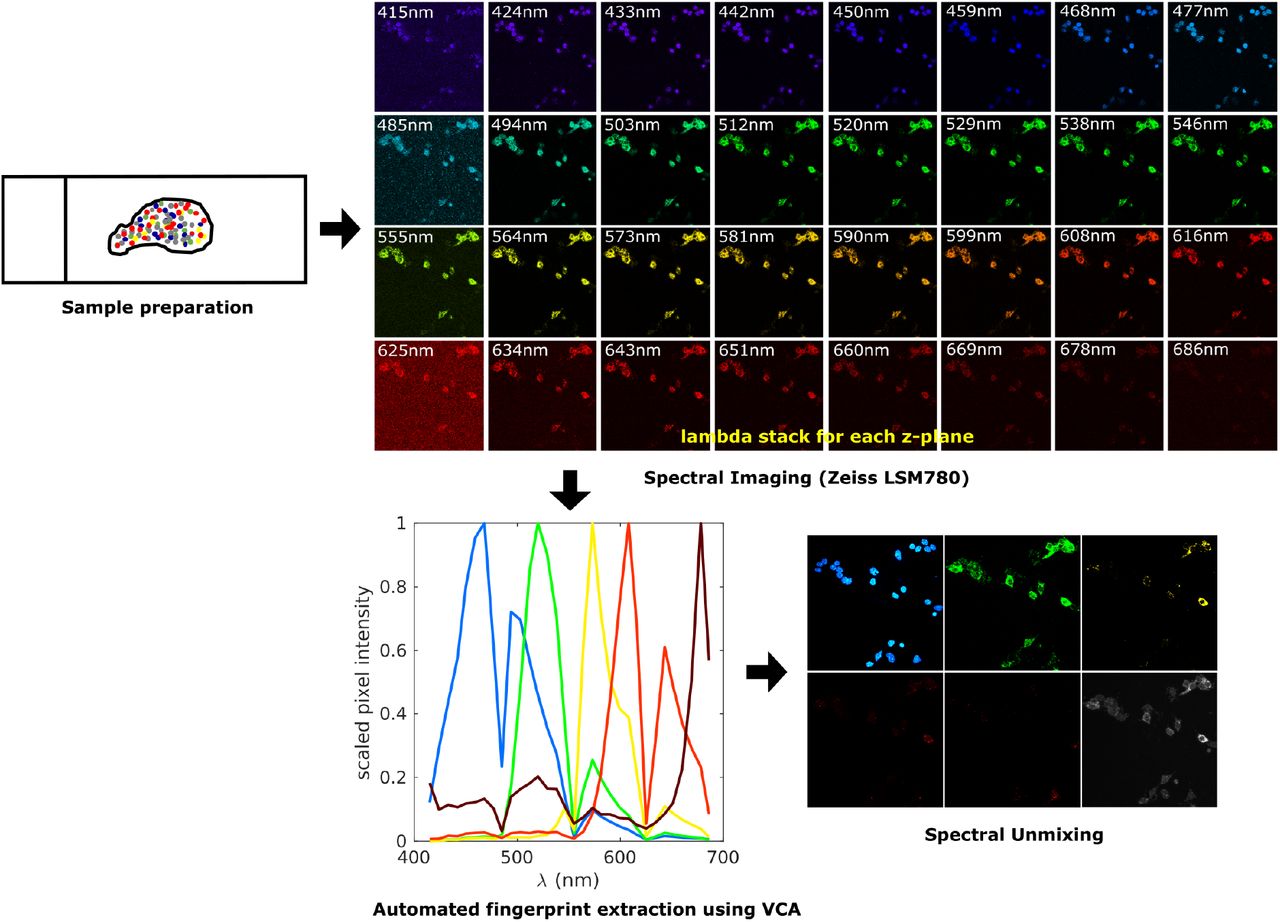
Vijay Sadashivaiah, Madhavi Tippani, Stephanie C Page, Sang Ho Kwon, Svitlana V Bach, Rahul A Bharadwaj, Thomas M Hyde, Joel E Kleinman, Andrew E Jaffe, Kristen R Maynard, BMC Neuroscience, 2021 paper / code An automated approach to spectral unmixing of fluorescent images |
|
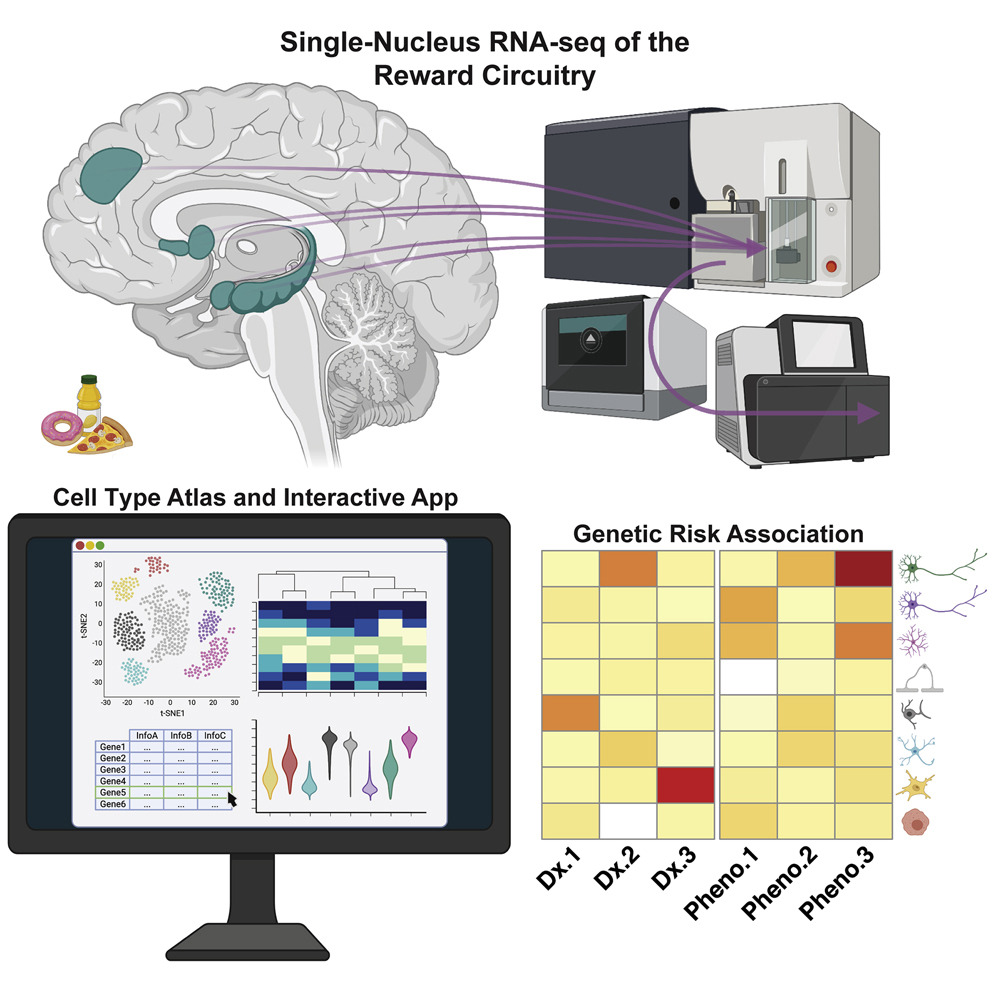
Matthew N Tran, Kristen R Maynard, Abby Spangler, Louise A Huuki, Kelsey D Montgomery, Vijay Sadashivaiah, Madhavi Tippani, Brianna K Barry, Dana B Hancock, Stephanie C Hicks, Joel E Kleinman, Thomas M Hyde, Leonardo Collado-Torres, Andrew E Jaffe, Keri Martinowich Neuron 2021 paper / code A single-nucleus RNA-sequencing resource of 70,615 high-quality nuclei to generate a molecular taxonomy of cell types across five human brain regions. |
|
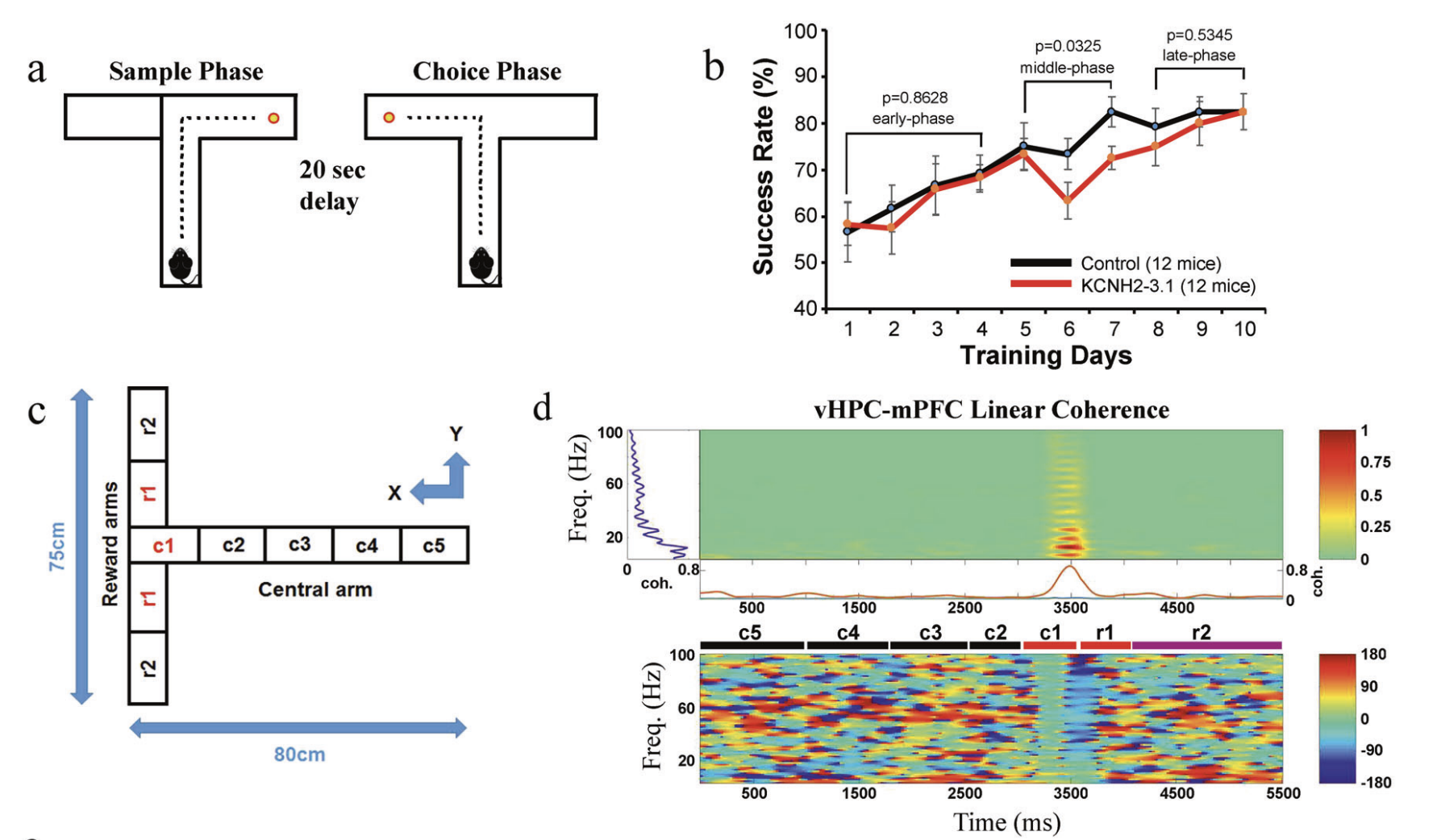
Ming Ren, Zhonghua Hu, Dr. Qiang Chen, Andrew E Jaffe, Yingbo Li, Vijay Sadashivaiah, Shujuan Zhu, Nina Rajpurohit, Joo Heon Shin, Wei Xia, Yankai Jia, Jingxian Wu, Sunny Lang Qin, Xinjian Li, Jian Zhu, Qingjun Tian, Daniel Paredes, Fengyu Zhang, Kuan Hong Wang, Venkata S Mattay, Joseph H Callicott, Karen F Berman, Daniel R Weinberger, Feng Yang Molecular Psychiatry 2019 paper |
|
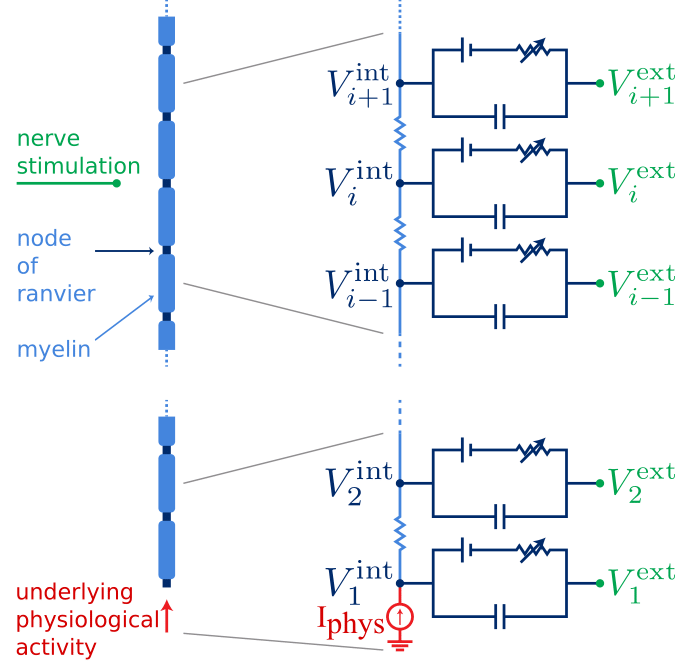
Vijay Sadashivaiah, Pierre Sacré, Yun Guan, William S Anderson, Sridevi V Sarma Journal of Computational Neuroscience 2019, EMBC 2018, EMBC 2017 paper 1 / paper 2 / paper 3 / paper 4 / code We constructed a mechanistic, stochastic and functional models of nerve fiber to quantify interactions. |
|
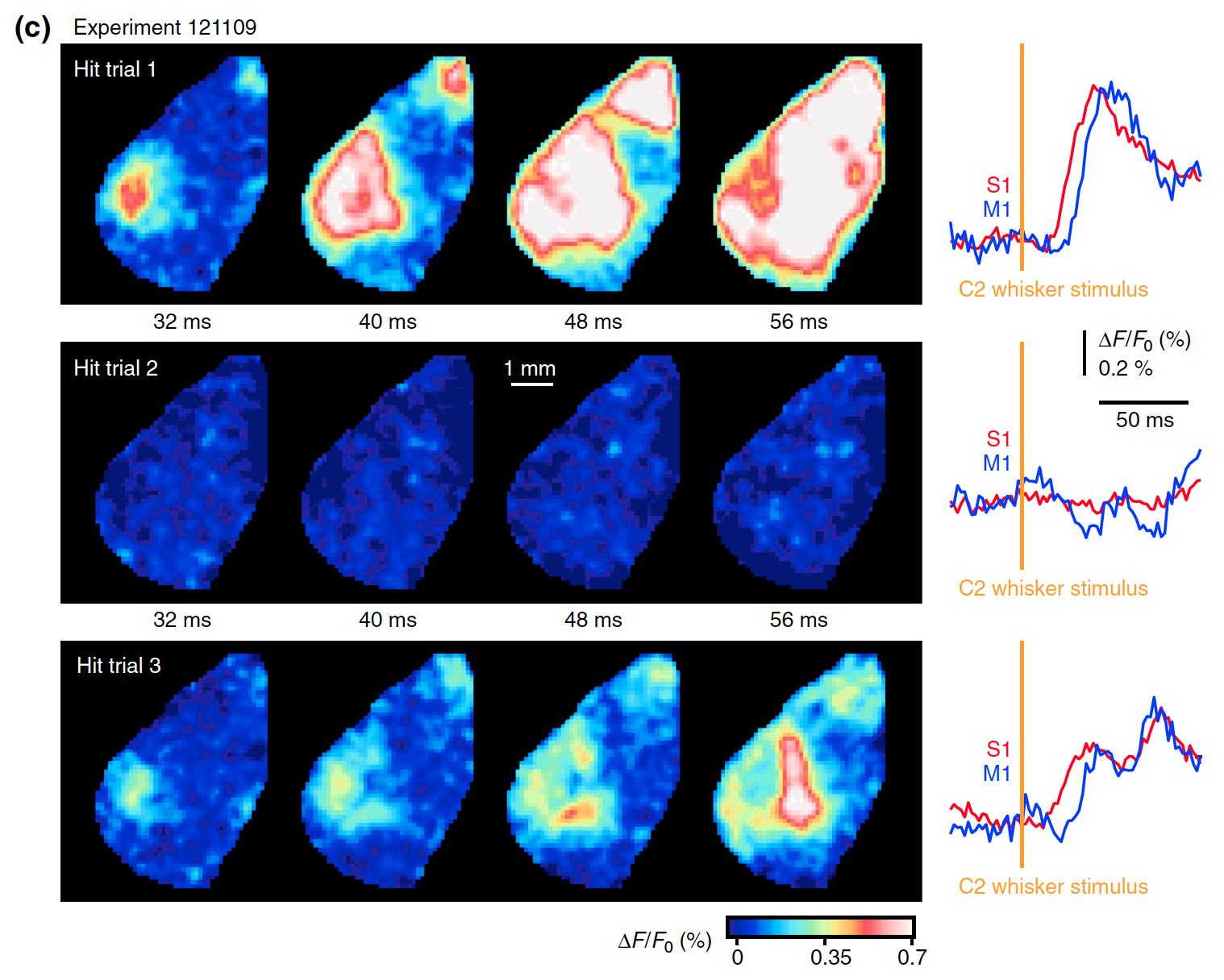
Alexandros Kyriakatos, Vijay Sadashivaiah, Yifei Zhang, Alessandro Motta, Mattieu Auffret, Carl CH Petersen Neurophotonics 2017 paper We studied the sensory motor interactions in mice brain using voltage sensitive dye imaging. |
|
© 2025 Vijay Sadashivaiah | Source code credit to Dr. Jon Barron |